ViewerImageProxy: allow data operations on plotted mosaics#812
ViewerImageProxy: allow data operations on plotted mosaics#812ejeschke merged 20 commits intoejeschke:masterfrom
Conversation
|
@pllim, this is preliminary WIP, but far enough along now for testing and some feedback. Hold the code review for now, as there is much to do there, but try out the In a nutshell, what this does is replace the idea of building one large image with all the tiles in it, and instead just plots them on the ginga canvas relative to the WCS. Advantages are that saves memory, can be faster to create the mosaic and doesn't require the user to specify a FOV size. Indeed, the tiles can be quite far apart, as they are simply not plotted unless in the clip area. The downside: most Ginga plugins have been written with the idea that there is a single image "loaded" into the viewer. Many of them work by trying to get that single image and do something with it. This PR makes a kind of "proxy" that supports some of the image functions; in particular, the functions meant for grabbing small subsets of data and doing some analysis on that. This PR has been written to allow them to make only minor modifications to work with both "regular" images and the mosaics. |
0e6e9da to
fe64f6d
Compare
This comment has been minimized.
This comment has been minimized.
This comment has been minimized.
This comment has been minimized.
bb49617 to
82c357a
Compare
This comment has been minimized.
This comment has been minimized.
This comment has been minimized.
This comment has been minimized.
This comment has been minimized.
This comment has been minimized.
1173f90 to
0395b84
Compare
This comment has been minimized.
This comment has been minimized.
Ping! 😺 |
This comment has been minimized.
This comment has been minimized.
0395b84 to
6bd4f4b
Compare
|
Rebased. |
I guess I'll have to go into https://github.com/spacetelescope/stginga/blob/master/stginga/plugins/MosaicAuto.py and make it check for Ginga version and apply the correct API, but since you said this is WIP and not ready for code review, I'll hold off on that.
|
|
To clarify, for the
|
|
 pllim
left a comment
pllim
left a comment
There was a problem hiding this comment.
I posted my findings in the comments above.
I am also concerned about taking away the ability to save the output mosaic. Is there a way to retain the "classic" mosaic plugin so I can still save the mosaic out somehow?
Has it returned? Does it happen every time you start up ginga from terminal? |
We could create
This is kind of a limitation of this approach--because no single image is "built", there is no one large image that can be written out as a FITS file. |
I'm certainly not opposed to keeping the existing plugin around--we still have uses for that here at Subaru. I think we'd just need to be careful in naming and documentation, so that new users don't get confused between the two plugins. |
|
Existing vs. New plugin New plugin
Existing plugin
|
31fd259 to
daae783
Compare
Pushed a fix for this.
Can you show an example with FITS for this? I worked really hard to squash that one and was pretty sure I had figured it out. Did it happen when opening a file in the usual way or when using |
Not sure what happened, but it looks like it ran when I pushed this last fix. Seems ok. |
Any FITS file will do. I used an example HST/ACS image (name goes like
I only see ReadTheDocs reporting back. That one only builds the documentation for this PR. I do not see any result from TravisCI. Seems like the CI ran but not reporting back -- https://travis-ci.org/github/ejeschke/ginga/pull_requests |
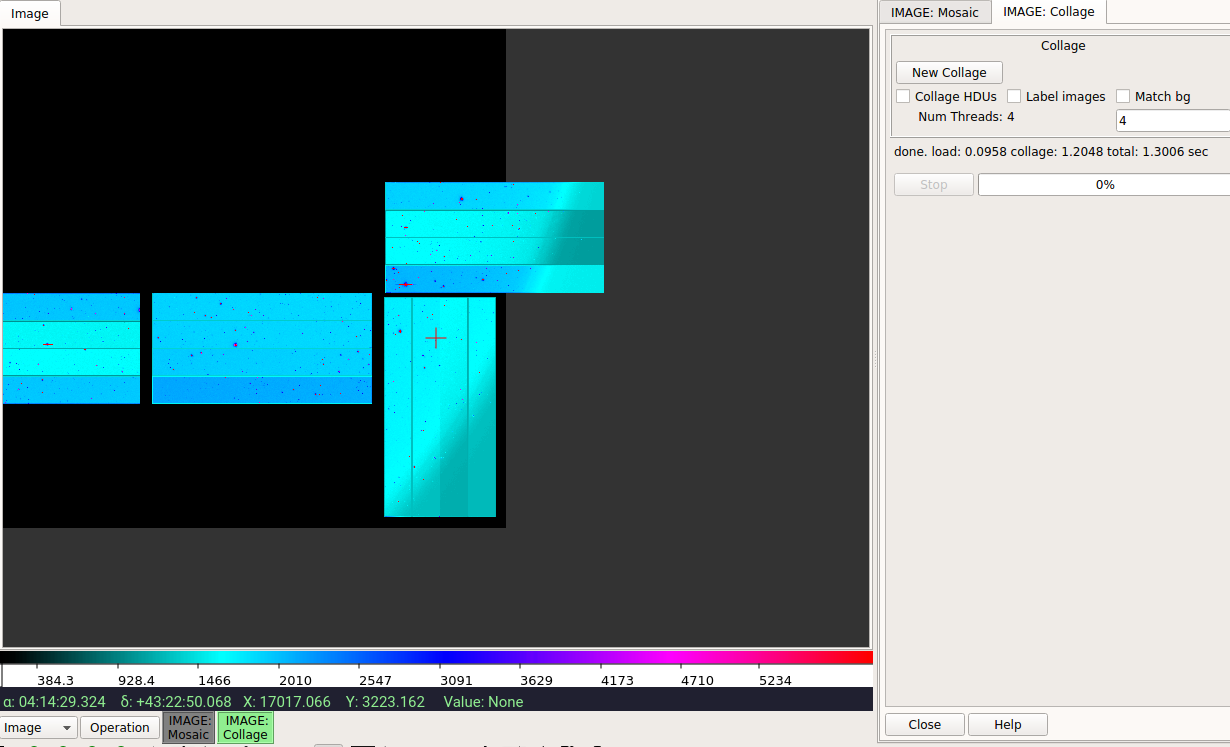
I assume you mean "buffer from Mosaic and then use Collage"? Unless you press "New Collage" after starting the Using I agree that these two plugins really don't play well together and the user should choose one or the other and not mix them. I can add some more documentation on |
Isn't this it? |
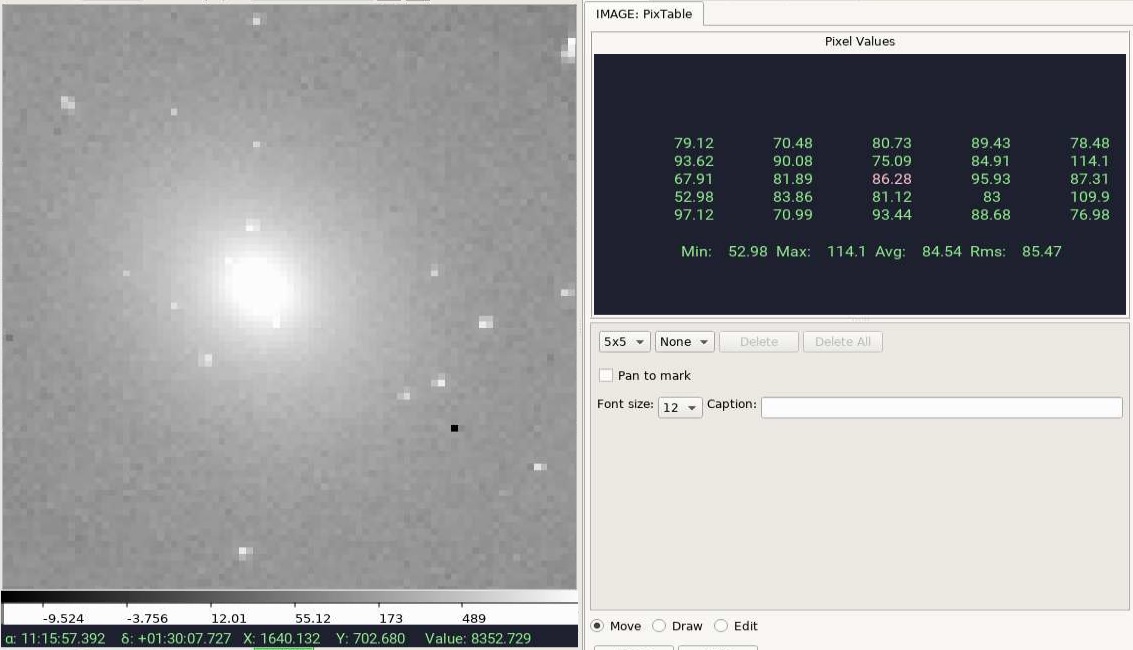
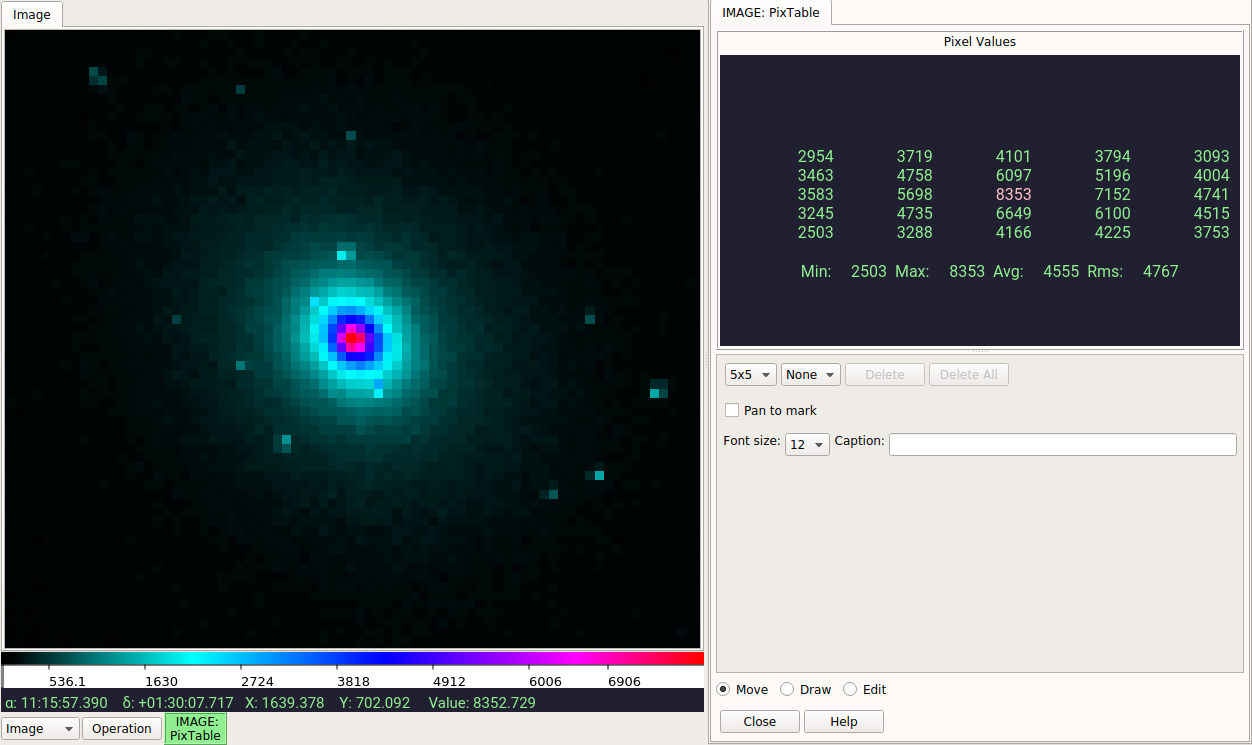
I just tried several j...fits images in my testdata directory and they are all working fine (image reported in Value matches highlighted value in |
I see what you mean now. Wonder what the heck is going on with that. I'll try rebasing another one of the PRs and see if the same thing happens. |
|
I just rebased the |
|
Maybe your Travis CI was still using GitHub Services? https://developer.github.com/v3/guides/replacing-github-services/ |
|
I suppose so. I don't remember touching that for a long time. |
|
Image link: https://stsci.box.com/s/3k009ufudki2c5lcug1qjwnscbpqw2wm What I see ( I am using Ginga via SSH, so I am not physically in front of the machine where it is running. Does this matter though? |
|
p.s. Yes, I picked up the latest changes from this PR. The |
|
Apparently, ginga should have been auto-migrated on February 1, 2019. I guess that didn't work... |
Not sure why you were getting a |
|
@pllim, I am seeing matching values. Your |
github shows a web hook registered for Travis. History of events shows the hooks being called. Travis builds show that the hooks are triggering builds. @pllim, is |
|
Re: |
|
Re: segfault -- Also on |
|
I think the blockers are all not related to this PR. Would still be nice to see Travis CI actually reporting back before merge but also not a blocker. |
You can see the Travis result passing here. Did |
|
I can't speak for
|
|
I also see it under |
So that is that is confusing, because this is the exact webhook that is shown in Ginga repo settings. I'm wondering if I also need to install the github app in order to get the result to show up in the checks tab. |
|
Well, doesn't hurt to try. The webhook is obviously sending jobs to Travis CI, just somehow not reporting back. |
|
Thanks for filing the new issues, @pllim. In the interest of working through the backlog of PRs, I'm going to merge this one. I will add some more documentation to clarify usage of |
|
Sounds good. Thanks! |

Adds a
ViewerImageProxyclass, which is a kind of stand-in for an image (i.e.BaseImage-based) wrapper. It looks at images plotted on the canvas and provides SOME methods to treat it as though it was one large single image. This allows us to save large amounts of memory for mosaiced fields,because we don't need to allocate one huge ndarray to hold everything, but only plot each image at the proper location and orientation on the viewer's canvas--like a sparse data array.
Several plugins are modified so that they will work with this kind of mosaiced display, particularly
Pick,Cuts,Histogram,PixTable,Pan.